SC02 Panelist: Computational Biology and High Performance Computing

Moderator: Craig Stewart, Indiana University
Panelists: Chris Johnson (Utah); John Reynders (Celera); David Bader (New Mexico); Debra Goldfarb (IDC); Rick Stevens (Argonne National Laboratories and the University of Chicago)
The biological/biomedical/bioinformatic/genomic/proteomic sciences now require the use of massive high performance computing (HPC) resources. In fact, biological sciences may more than double the size of the HPC market according to some estimates. This panel will be drawn primarily from the user community - people whose background is in the biological or chemical sciences - who have been successful in implementing HPC solutions to current biological and biomedical problems.
The questions to be addressed by this panel include the following:
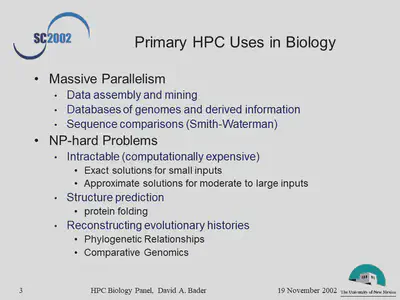
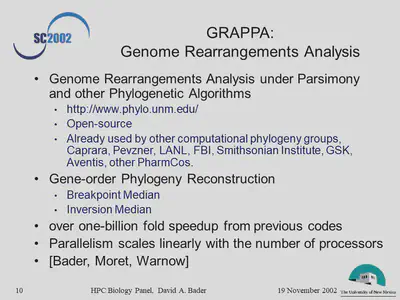
- What has been the nature of the problems and the HPC solutions when HPC technologies have been critical to the solution of important problems?
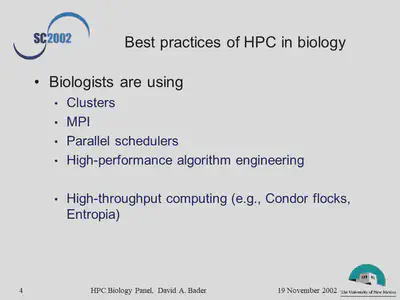
- What are the current best practices in use of HPC in biological/biomedical sciences?
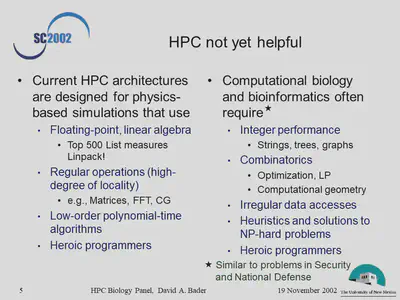
- Where has use of HPC approaches not yet been helpful in solving biological problems, and why not?
- What are the big future problems in the biological sciences that will require use of HPC systems?
- What does the biological research community need from the HPC community in order to foster better and more rapid implementation of HPC solutions to biological problems?
- What hardware architectures are most effective now, and what will the most effective hardware architectures be in 5 years?
- What are the software frameworks and applications that most effectively facilitate solution of important biological and biomedical problems?