David A. Bader
Distinguished Professor and Director of the Institute for Data Science
New Jersey Institute of Technology
Biography
David A. Bader is a Distinguished Professor and founder of the Department of Data Science in the Ying Wu College of Computing and Director of the Institute for Data Science at New Jersey Institute of Technology. Prior to this, he served as founding Professor and Chair of the School of Computational Science and Engineering, College of Computing, at Georgia Institute of Technology. Bader is an elected Board Member of the Computing Research Association (CRA). He is a Fellow of the IEEE, ACM, AAAS, and SIAM; a recipient of the IEEE Sidney Fernbach Award; the 2022 Innovation Hall of Fame inductee of the University of Maryland’s A. James Clark School of Engineering; a 2025 inductee of the Mimms Museum of Technology and Art’s Hall of Fame; and the 2025 recipient of the Heatherington Award for Technological Innovation. The Computer History Museum recognizes Bader for developing the first Linux-based supercomputer which became the predominant architecture for all major supercomputers in the world. In 2025, HPCwire named Bader as one of its “35 Legends”. In 2026, Scientific Computing World named Bader as one of its 25 “high-performance computing thought leaders in scientific computing”.
Interests
- Data Science
- High Performance Computing
- Real-World Analytics
Education
PhD in Electrical Engineering, 1996
University of Maryland
MS in Electrical Engineering, 1991
Lehigh University
BS in Computer Engineering, 1990
Lehigh University
Biography
Dr. Bader is a Fellow of the IEEE, ACM, AAAS, and SIAM; a recipient of the IEEE Sidney Fernbach Award; the 2022 Innovation Hall of Fame inductee of the University of Maryland’s A. James Clark School of Engineering; a 2025 inductee of the Mimms Museum of Technology and Art’s Hall of Fame; and the 2025 recipient of the Heatherington Award for Technological Innovation. He advises the White House, most recently on the National Strategic Computing Initiative (NSCI) and Future Advanced Computing Ecosystem (FACE). Bader is a leading expert in solving global grand challenges in science, engineering, computing, and data science. His interests are at the intersection of high-performance computing and real-world applications, including cybersecurity, massive-scale analytics, and computational genomics, and he has co-authored over 400 scholarly papers and has best paper awards from ISC, IEEE HPEC, and IEEE/ACM SC. Dr. Bader has served as a lead scientist in several DARPA programs including High Productivity Computing Systems (HPCS) with IBM, Ubiquitous High Performance Computing (UHPC) with NVIDIA, Anomaly Detection at Multiple Scales (ADAMS), Power Efficiency Revolution For Embedded Computing Technologies (PERFECT), Hierarchical Identify Verify Exploit (HIVE), and Software-Defined Hardware (SDH). Recently, Bader received an NVIDIA AI Lab (NVAIL) award, and a Facebook Research AI Hardware/Software Co-Design award.
Dr. Bader has previously served as the Editor-in-Chief of the ACM Transactions on Parallel Computing, and as Editor-in-Chief of the IEEE Transactions on Parallel and Distributed Systems. He serves on the leadership team of Northeast Big Data Innovation Hub as the inaugural chair of the Seed Fund Steering Committee. ROI-NJ recognized Bader as a technology influencer on its 2021 inaugural and 2022 lists. In 2012, Bader was the inaugural recipient of University of Maryland’s Electrical and Computer Engineering Distinguished Alumni Award. In 2014, Bader received the Outstanding Senior Faculty Research Award from Georgia Tech. Bader is a member of Tau Beta Pi (National Engineering Honor Society), Eta Kappa Nu (Electrical Engineering Honor Society), and Omicron Delta Kappa (National Leadership Honor Society). Bader has also served as Director of the Sony-Toshiba-IBM Center of Competence for the Cell Broadband Engine Processor and Director of an NVIDIA GPU Center of Excellence. In 1998, Bader built the first Linux supercomputer that led to a high-performance computing (HPC) revolution, and Hyperion Research estimates that the total economic value of Linux supercomputing pioneered by Bader has been over $100 trillion since its inception. The Computer History Museum recognizes Bader for developing the first Linux-based supercomputer which became the predominant architecture for all major supercomputers in the world. Bader is a cofounder of the Graph500 List for benchmarking “Big Data” computing platforms. He is recognized as a “RockStar” of High Performance Computing by InsideHPC and as HPCwire’s People to Watch in 2012 and 2014. In 2025, HPCwire named Bader as one of its “35 Legends”. In 2026, Scientific Computing World named Bader as one of its 25 “high-performance computing thought leaders in scientific computing”.
Media Appearances

Experience
Recent Boards
Recent Posts
/Unsplash]](/post/20260508-fastcompany/p-1-91538458-google-used-to-be-a-search-engine-now-it-wants-to-be-everything_hu_a58c6cf0ba56d32f.webp)
Twenty years ago, if you asked the average person what Google was, they’d tell you it was a search engine. The company became synonymous with searching for information online, reaching a level of dominance no search engine had seen before, or has seen since.
Projects
This award supports the development of advanced computational methods for tracking and analyzing evolving patterns in large-scale networks. Patterns of connections among entities, known as subgraphs, underpin insights in domains such as social interactions, biological processes, financial transactions, and communication systems. Real-time analysis of how these patterns form and dissolve can enable early detection of disease outbreaks, improved understanding of social dynamics, and enhanced network security. By creating scalable and accessible tools for dynamic network analysis, this project will advance the national interest in data-driven discovery across science, technology, and public welfare.
Community detection methods enable an understanding of the structure of networks at multiple scales. While many methods exist, only a few are able to scale to large networks and/or are implemented in large computational infrastructure. As we have recently shown, even those that scale to large datasets, fail to reliably produce well-connected clusters. Finally, given that the choice of clustering method depends on both the network being analyzed and the question of interest, providing the domain specialist a choice of multiple clustering methodologies within a common framework for exploratory data analysis, is essential. This project will make substantial advances on these challenges through the coordinated development of advanced cyber-infrastructure, scalable to very large networks, that offers multiple options for community detection, search, and extraction. The infrastructure will be accessible across platforms ranging from laptops to multi-node clusters with distributed memory.
A real-world challenge in data science is to develop interactive methods for quickly analyzing new and novel data sets that are potentially of massive scale. This award will design and implement fundamental algorithms for high performance computing solutions that enable the interactive large-scale data analysis of massive data sets. Based on the widely-used data types and structures of strings, sets, matrices and graphs, this methodology will produce efficient and scalable software for three classes of fundamental algorithms that will drastically improve the performance on a wide range of real-world queries or directly realize frequent queries. These innovations will allow the broad community to move massive-scale data exploration from time-consuming batch processing to interactive analyses that give a data analyst the ability to comprehensively, deeply and efficiently explore the insights and science in real world data sets. By enabling the increasing number of developers to easily manipulate large data sets, this will greatly enlarge the data science community and find much broader use in new communities. Materials from this project will be included in graduate and undergraduate course curriculum. Especially, women, high school students and other underrepresented groups in STEM areas will be encouraged to participate in this research activity.
Research Directions
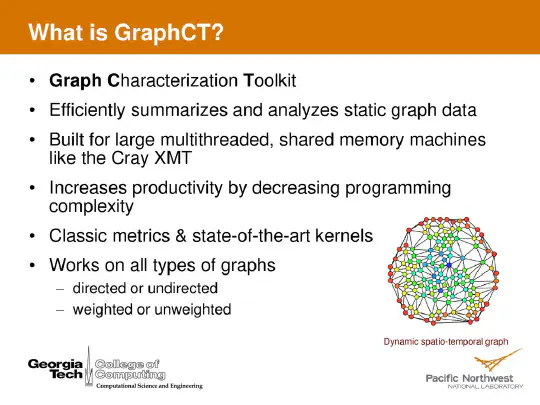
Graph algorithms represent some of the most challenging known problems in computer science for modern processors. These algorithms contain far more memory access per unit of computation than traditional scientific computing. Access patterns are not known until execution time and are heavily dependent on the input data set. Graph algorithms vary widely in the volume of spatial and temporal locality that is usable my modern architectures. In today’s rapidly evolving world, graph algorithms are used to make sense of large volumes of data from news reports, distributed sensors, and lab test equipment, among other sources connected to worldwide networks. As data is created and collected, dynamic graph algorithms make it possible to compute highly specialized and complex relationship metrics over the entire web of data in near-real time, reducing the latency between data collection and the capability to take action.
Facebook AI Systems Hardware/Software Co-Design research award on Scalable Graph Learning Algorithms
https://research.fb.com/blog/2019/05/announcing-the-winners-of-the-ai-system-hardware-software-co-design-research-awards/
Deep learning has boosted the machine learning field at large and created significant increases in the performance of tasks including speech recognition, image classification, object detection, and recommendation. It has opened the door to complex tasks, such as self-driving and super-human image recognition. However, the important techniques used in deep learning, e.g. convolutional neural networks, are designed for Euclidean data type and do not directly apply on graphs. This problem is solved by embedding graphs into a lower dimensional Euclidean space, generating a regular structure. There is also prior work on applying convolutions directly on graphs and using sampling to choose neighbor elements. Systems that use this technique are called graph convolution networks (GCNs). GCNs have proven to be successful at graph learning tasks like link prediction and graph classification. Recent work has pushed the scale of GCNs to billions of edges but significant work remains to extend learned graph systems beyond recommendation systems with specific structure and to support big data models such as streaming graphs.
Dynamic graphs are all around us. Social networks containing interpersonal relationships and communication patterns. Information on the Internet, Wikipedia, and other datasources. Disease spread networks and bioinformatics problems. Business intelligence and consumer behavior. The right software can help to understand the structure and membership of these networks and many others as they change at speeds of thousands to millions of updates per second.
Books
Expertise in massive scale graph analytics is key for solving real-world grand challenges from health to sustainability to detecting insider threats, cyber defense, and more. Massive Graph Analytics provides a comprehensive introduction to massive graph analytics, featuring contributions from thought leaders across academia, industry, and government.
The hybrid/heterogeneous nature of future microprocessors and large high-performance computing systems will result in a reliance on two major types of components: multicore/manycore central processing units and special purpose hardware/massively parallel accelerators. While these technologies have numerous benefits, they also pose substantial performance challenges for developers, including scalability, software tuning, and programming issues.
Although the highly anticipated petascale computers of the near future will perform at an order of magnitude faster than today’s quickest supercomputer, the scaling up of algorithms and applications for this class of computers remains a tough challenge. From scalable algorithm design for massive concurrency toperformance analyses and scientific visualization, Petascale Computing: Algorithms and Applications captures the state of the art in high-performance computing algorithms and applications. Featuring contributions from the world’s leading experts in computational science, this edited collection explores the use of petascale computers for solving the most difficult scientific and engineering problems of the current century.
Featured Publications
Recent Publications
Recent & Upcoming Talks
Session Topics
- Is academia for me?
- How do I land an academic position?
About of Featured Guests:
- David Bader (ECE Ph.D. ‘96), Professor, New Jersey Institute of Technology
- Chai-Wai Wong (ECE Ph.D. ‘17), Associate Professor, NC State University
Local Panelists:
- Prof. Sanghamitra Dutta
- Prof. Carol Espy-Wilson
- Prof. Jesse Moody
- Prof. Thomas Murphy
- Prof. Kaiqing Zhang
Moderator:
- Prof. Cheng Gong
Please register at https://go.umd.edu/ECEGSA-AcademicPanel2025
Event counts as one ECE Colloquium Series attendance
Event organizers: Irem Didin, Sydney Overton and Solaleh Mohammadi
Contact
- Institute for Data Science, New Jersey Institute of Technology, 101 Hudson St., Suite 3610, Jersey City, NJ 07302
- DM Me
- Chat on Keybase